| EECS 311: Graphs |
Home Course Info Links Grades |
Lectures Newsgroup Homework Exams |
There's a lot of terminology associated with graphs, much of it similar to that used in trees. Trees are actually a special case of graphs, but they're the most useful special case.
You should be familiar with all of the following:
This is the simplest, most flexible method. Given a graph like this

you create a list of lists, where each list starts with a node, followed by that node's successors:
((a (b 1) (d 5)) (b (a 1) (c 1)) (c (b 3) (d 1)) (d (b 2)) )
To find a node, you search the list of lists sequentially, looking at the first element in each list, until you find the desired node.
Almost the same, but you have an array of pointers to lists of successors, indexed by nodes.
This is obviously faster to get to a particular node, but only works for node identifiers that come from a reasonable enumerable set, like characters. It wouldn't work well for cities in the United States.
This is also quite simple. Given N nodes, from an enumerable set, just make an NxN matrix Adj. The value in Adj[i, j] says whether j is an immediate successor of i or not.
For our graph above, it would be
| a | b | c | d | |
|---|---|---|---|---|
| a | 0 | 1 | - | 5 |
| b | 1 | 0 | 1 | - |
| c | - | 3 | 0 | 1 |
| d | - | 2 | - | 0 |
Consider prerequisites for courses, such as those for computer engineering at Northwestern.
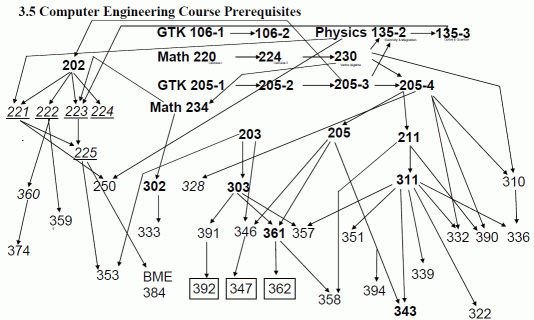
This is a directed acyclic graph.
One of the things you want to know is what order courses can be taken in. You couldn't, for example, take 374 before 202, because 374 requires 360, 360 requires 222, and 222 requires 202. But you could take 374 before or after 359.
A topological ordering (or sort or linearization) is any sequence of the nodes (vertices) in the graph, such that, if node2 follows node1 in some path in the graph, then node2 must come after node1 in the topological ordering. A graph may have many valid topological orderings. (A graph with cycles has none.)
Here's the standard algorithm for a topological sort, given a graph with N nodes:
indegree[N],
where indegree[i] represents how
many nodes point directly to nodei.indegree[i]
is 0.indegree[j] for
every nodej that X points toindegree[j] is now 0, enqueue nodejWhat's the complexity of this algorithm?
Given two nodes in a graph, a common problem is to find a path from one to the other.
This method finds a path, pretty cheaply, but may not find the shortest path. It uses a stack. The gist of the algorithm is this:
To make this algorithm useful, we need to keep track of the path we found. To do that, we actually need several stacks and the algorithm will look very much like our course scheduler algorithm, keeping track of all alternatives and what's been tried so far.
Also, if the graph has cycles or nodes that many other nodes link to, it will loop or at least be very inefficient.
This is easy to fix by keeping track of which nodes have been visited and never following the successors of any already visited node.
This method finds the shortest path, but it uses more space to do so. It uses a queue. The gist of the algorithm is this:
Again, to make this algorithm useful and practical, we need to keep track of the path and which nodes have been visited. You should be able to figure out why this algorithm will not loop endlessly if a path exists, even if there are cycles in the graph, even without tracking visited nodes. But to avoid endless loops in the "no path" case, and for efficiency, we should track visited nodes.
If we have an unweighted graph, then the shortest path is the cheapest one and we can use breadth-first search to find it.
If we have weighted edges, however, then there may be a path with many links that costs less than a path with fewer links.
Finding lowest cost paths is a core problem in many applications, from finding the cheapest or fastest airplane route to somewhere to finding the least loaded set of telephone connections between two points.
The obvious solution is to modify the depth-first traversal algorithm to:
This will try many paths. For example, for the network in Figure 9.15, it will try
and I may have missed some.
Obviously, many of these paths include the same subpaths over and over. If we kept track of those subpaths, we'd reduce our work significantly.
Dijsktra's algorithm incrementally grows the set of cheapest paths from a starting node, start, breadth-first, until it reaches a given destination node, end.
It uses the following data structures in addition to Adj[i, j]:
The algorithm looks like this:
unprocessed = { <start, 0> } cost[i] = empty While unprocessed is not empty Pop <i, cost> off unprocessed. If i = end, exit with <i, cost>. If cost[i] is unknown cost[i] = cost. For each node j with an edge to i push <j, cost[i] + Adj[i, j]> onto unprocessed
This is an O(N2) algorithm, in that we have to process O(N) nodes, and for each node, it takes O(N) steps to find i and O(N) steps to update the costs for each node j.
Try running Dijkstra's algorithm on the graph given at the start of this page.
Note that this algorithm doesn't tell you the shortest path. What would you add to the above to keep track of the best path? Hint: you could augment what you store in unprocessed and cost[].
Also note that we can avoid pushing <j, cost> onto unprocessed in two cases:
These don't change the O() complexity. The O(log n) check for cost[j] might even be slower than the O(log n) time to add <j, cost> to unprocessed. This is an empirical issue.
Dijskstra's algorithm is fine for finding the cheapest path between given pair of nodes. If we wanted to find all the cheapest paths between every pair of nodes, there are O(N2) pairs of nodes, so we'd have an O(N4) algorithm if we simply called Dijskstra's algorithm on every pair.
Fortunately, there's a simpler cheaper way. If we know we're doing all N2 pairs, i.e., we're making a 2-dimensional table Cost of shortest paths, then we use Floyd's algorithm.
Cost[i, j] = Adj[i, j] for all i, j For each node k from 1 to N For each node i from 1 to N For each node j from 1 to N connect i to j via k if that path is shorter than the current path from i to j Cost[i, j] = min(Cost[i, j], Cost[i, k] + Cost[k, j])
That's it. The inner formula says that if the path i - k - j is cheaper than the previously known path (which, because of the way we're generating them, could not have included k), make that the new cost of getting from i to j.
This is an O(N3) algorithm, because of the three nested O(N) loops. The correctness of this simple algorithm is not immediately obvious. A good discussion of why it works is here.
For example, when we consider node A, we calculate costs for paths through A for all nodes that we know can get to A, say B and C. Then we do B, C, and so on. Suppose we later find out that node G can get to A. Won't we have missed a possible short path from G to B or C? Read the proof.
Notice that this version of the algorithm doesn't actually create the paths. Nowhere does it store what the paths are from i to k, k to j, or i to j, just what the cost of those paths are. To do so, we'd need to use a 2D Paths array, and add the following line to the innermost loop above:
Paths[i, j] = k if Cost[i, k] + Cost [k, j] < Cost [i, j]
Try running Floyd's algorithm, with Paths, on the graph given at the start of this page.
For unweighted graphs, Warshall's algorithm can be used to convert an adjacency matrix into a connectivity matrix. That is, we start with a matrix of 1's and 0's indicating what nodes are adjacent to what, run the algorithm, and end up with a matrix of 1's and 0's indicating what nodes are can be reached from what nodes. A 1 in [i, j] means that there's some path from i to j.
Not surprisingly, the algorithm is very similar to Floyd's algorithm, with a simpler function for changing the array of connections.
Initialize: Conn[i, j] = Adj[i, j] for all i, j For each node k from 1 to N For each node i from 1 to N For each node j from 1 to N Connect i and j if both connect to k Conn[i, j] = Conn[i, j] | ( Conn[i, k] & Conn[k, j] )
This is an O(N3) algorithm, because of the three nested O(N) loops.
A graph is not a tree, but we can remove edges until we have a tree that still connects all vertices. If the edges of the graph have costs, a minimum spanning tree (MST) is such a tree with the lowest cost. MST's are useful because, being trees, they are much easier to work with than graphs, but they often capture the critical information we need from a graph.
Prim's algorithm for constructing an MST is similar to Dijkstra's algorithm. It starts from some node and adds edges in a greedy fashion (where greedy means picking the edge that leads to the least cost increase).
Initialize the tree with a lowest cost edge. While there are nodes not connected to the tree Add the lowest cost edge from a node in the tree to a node not in the tree.
Kruskal's algorithm is also a greedy algorithm, but it collects the lowest cost edges that don't cause cycles, rather than growing a single tree. The algorithm can be implemented using priority queues and union/find.
Let MST be an empty set of edges. Let PQ be a priority queue of all edges, with the lowest cost on top. Put each node in a separate set. While there is more than one set Take the top edge (u, v) from PQ. If find(u) != find(v), add the edge to MST union find(u) with find(v).
Comments?  Send mail to
c-riesbeck@northwestern.edu.
Send mail to
c-riesbeck@northwestern.edu.